中国临床药理学与治疗学 ›› 2026, Vol. 31 ›› Issue (3): 289-299.doi: 10.12092/j.issn.1009-2501.2026.03.001
郑凯乐1,2,3(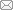 ), 付嘉钊4, 尤佳1,2,3, 吴丹5, 王学彬2,3,*(
), 付嘉钊4, 尤佳1,2,3, 吴丹5, 王学彬2,3,*( ), 王卓3
), 王卓3
收稿日期:2025-01-07
修回日期:2025-10-14
出版日期:2026-03-26
发布日期:2026-04-03
通讯作者:
王学彬
E-mail:1604751742@qq.com;binxuewang@sjtu.edu.cn
作者简介:郑凯乐,女,硕士研究生,药师,研究方向:免疫抑制剂个体化给药。E-mail:基金资助:
Kaile ZHENG1,2,3( ), Jiazhao FU4, Jia YOU1,2,3, Dan WU5, Xuebin WANG2,3,*(
), Jiazhao FU4, Jia YOU1,2,3, Dan WU5, Xuebin WANG2,3,*( ), Zhuo WANG3
), Zhuo WANG3
Received:2025-01-07
Revised:2025-10-14
Online:2026-03-26
Published:2026-04-03
Contact:
Xuebin WANG
E-mail:1604751742@qq.com;binxuewang@sjtu.edu.cn
摘要:
目的: 揭示肾移植受者术后发生抗体介导排斥反应(antibody-mediated rejection,ABMR)的潜在机制。方法: 选取肾移植术后 6 个月以上的受者作为研究对象,利用单细胞RNA测序(single-cell RNA sequencing,scRNA-seq)技术,对受者外周血单个核细胞(peripheral blood mononuclear cell,PBMC)中的免疫细胞亚群展开高分辨率的scRNA-seq分析。采用流式细胞技术检测不同免疫细胞群中差异基因对ABMR 的影响。结果: 本研究通过单细胞测序技术分析了ABMR受者与正常对照组的差异基因表达,发现CD69、CD83、CD52、CD74和CX3CR1为主要的差异基因。在经典单核细胞中,ABMR受者的CD83和CD52基因表达量较正常组有所增加,而治疗后呈现降低趋势;在初始CD4+T细胞中,ABMR受者的CD69基因表达量较正常组显著下降;在NK细胞中,ABMR受者的CX3CR1基因表达量较正常组降低,而治疗后CX3CR1的几何平均值进一步降低,CD74的几何平均值则有增加趋势。结论: 经典单核细胞中 CD83 和 CD52 基因表达上调,与抗原提呈细胞的功能增强相关,从而促进 ABMR 的发生。初始 CD4+T 细胞中 CD69 基因表达上调,与 T 细胞的激活和增殖相关,进一步参与ABMR 的进程。相反,NK 细胞中 CX3CR1 基因表达下调,与 NK 细胞的免疫监视功能障碍相关,从而在一定程度上抑制 ABMR 的发生。
中图分类号:
郑凯乐, 付嘉钊, 尤佳, 吴丹, 王学彬, 王卓. 单细胞测序揭示肾移植受者抗体介导排斥反应的潜在机制[J]. 中国临床药理学与治疗学, 2026, 31(3): 289-299.
Kaile ZHENG, Jiazhao FU, Jia YOU, Dan WU, Xuebin WANG, Zhuo WANG. Single-cell sequencing reveals the underlying mechanism of antibody-mediated rejection in renal transplant recipients[J]. Chinese Journal of Clinical Pharmacology and Therapeutics, 2026, 31(3): 289-299.
| Cell type | Marker genes |
| T cell | CD3E, CD3D, CD3G |
| B cell | CD79B, CD19, CD79A |
| Fibroblast | COL1A1, COL1A2, COL3A1, POSTN |
| Epithelial cell | EPCAM, CDH1, KRT18, KRT8 |
| Neutrophil | Ly6g, Ncf1, Csf3r, Cd177, Sorl1 |
| Macrophage | CD14, CD68, CD163, CD209, AIF1 |
| Dendritic cell | Flt3, CD1C |
| Natural killer cell | Nkg7, Ifng, Klrd1, Klrc1, CD56 |
| Endothelial cell | VWF, CDH5, Pecam1 |
| Basophil | TPSB2, TPSAB1, TPSAB2, CPA3 |
| Plasma cell | CD27, CD38, XBP1, JCHAIN |
| Monocyte | CD14, CD300E, CD244 |
| CD8+NKT-like cell | CD3E, CD8A, CD161, KLRC1 |
| Classical monocyte | CD14, LYZ |
| Na?ve B cell | CD19, CD20, CD79A, IGLL1 |
| Na?ve CD4+T cell | CD3E, CD4, CD45RA, LEF1 |
| Na?ve CD8+T cell | CD3E, CD8A, CD45RA, LEF1 |
| Non-classical monocyte | CD14, CD16, FCGR3A |
| Plasmacytoid dendritic cell | CD123, CD303, IRF7, LY6E |
| Platelet | CD41, CD61, PF4, GPIBα |
| Pro-B cell | CD19, CD34, CD79A, IGLL1, RAG1 |
表 1
Table 1 Marking genes used to identify major cell types of immunity
| Cell type | Marker genes |
| T cell | CD3E, CD3D, CD3G |
| B cell | CD79B, CD19, CD79A |
| Fibroblast | COL1A1, COL1A2, COL3A1, POSTN |
| Epithelial cell | EPCAM, CDH1, KRT18, KRT8 |
| Neutrophil | Ly6g, Ncf1, Csf3r, Cd177, Sorl1 |
| Macrophage | CD14, CD68, CD163, CD209, AIF1 |
| Dendritic cell | Flt3, CD1C |
| Natural killer cell | Nkg7, Ifng, Klrd1, Klrc1, CD56 |
| Endothelial cell | VWF, CDH5, Pecam1 |
| Basophil | TPSB2, TPSAB1, TPSAB2, CPA3 |
| Plasma cell | CD27, CD38, XBP1, JCHAIN |
| Monocyte | CD14, CD300E, CD244 |
| CD8+NKT-like cell | CD3E, CD8A, CD161, KLRC1 |
| Classical monocyte | CD14, LYZ |
| Na?ve B cell | CD19, CD20, CD79A, IGLL1 |
| Na?ve CD4+T cell | CD3E, CD4, CD45RA, LEF1 |
| Na?ve CD8+T cell | CD3E, CD8A, CD45RA, LEF1 |
| Non-classical monocyte | CD14, CD16, FCGR3A |
| Plasmacytoid dendritic cell | CD123, CD303, IRF7, LY6E |
| Platelet | CD41, CD61, PF4, GPIBα |
| Pro-B cell | CD19, CD34, CD79A, IGLL1, RAG1 |
| Variable | Recipient 1 | Recipient 2 | Recipient 3 | Recipient 4 |
| Gender | Male | Female | Male | Male |
| Age | 15 | 48 | 54 | 51 |
| Weight (kg) | 41 | 58.1 | 68.5 | 85 |
| Clinical diagnosis | AR | ABMR | IgAN | Normal |
| Organ transplantation | DCD | DCD | DCD | DCD |
| Immune induction | ATG | basiliximab | ATG | ATG |
| Immunosuppressant | TAC+MPA | TAC+MPA+Pred | TAC+MPA+Pred | TAC+MPA+Pred |
| Tacrolimus C0 (ng/mL) | 4.5 | 11.1 | 2.1 | 7.9 |
| Treatment adjustment | Rituximab | Bortezomib | No adjustment | No adjustment |
| Serum creatinine (μmoI/L) | 144 | 172 | 680 | 89 |
| Uric acid (μmol/L) | 593 | 345 | 303 | 498 |
| Urea (mmol/L) | 15.9 | 16.1 | 14.7 | 4.7 |
| Albumin/globulin | 139 | 1.18 | 1.19 | 1.54 |
| Clinical manifestation | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure |
表 2
Table 2 Clinical basic data of 4 kidney transplant recipients (n=4)
| Variable | Recipient 1 | Recipient 2 | Recipient 3 | Recipient 4 |
| Gender | Male | Female | Male | Male |
| Age | 15 | 48 | 54 | 51 |
| Weight (kg) | 41 | 58.1 | 68.5 | 85 |
| Clinical diagnosis | AR | ABMR | IgAN | Normal |
| Organ transplantation | DCD | DCD | DCD | DCD |
| Immune induction | ATG | basiliximab | ATG | ATG |
| Immunosuppressant | TAC+MPA | TAC+MPA+Pred | TAC+MPA+Pred | TAC+MPA+Pred |
| Tacrolimus C0 (ng/mL) | 4.5 | 11.1 | 2.1 | 7.9 |
| Treatment adjustment | Rituximab | Bortezomib | No adjustment | No adjustment |
| Serum creatinine (μmoI/L) | 144 | 172 | 680 | 89 |
| Uric acid (μmol/L) | 593 | 345 | 303 | 498 |
| Urea (mmol/L) | 15.9 | 16.1 | 14.7 | 4.7 |
| Albumin/globulin | 139 | 1.18 | 1.19 | 1.54 |
| Clinical manifestation | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure | Normal body temperature, stable blood pressure |
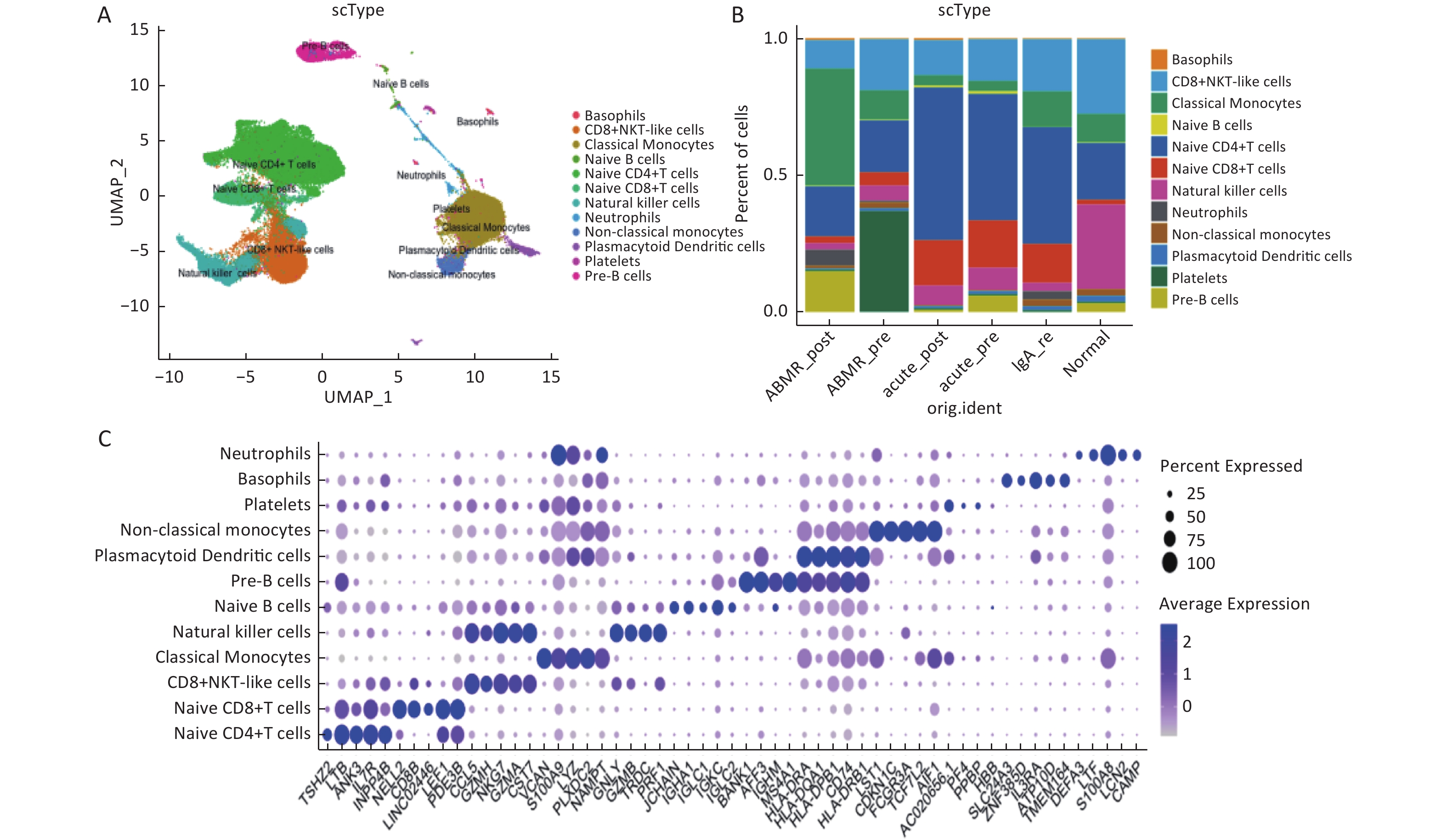
图 1
Fig.1 Immune cell subsets in peripheral blood of ABMR kidney transplant recipients A: UAMP plots of 12 major cell types; B: proportion of major cell types in different samples; C: expression levels of marker genes in major cell types.
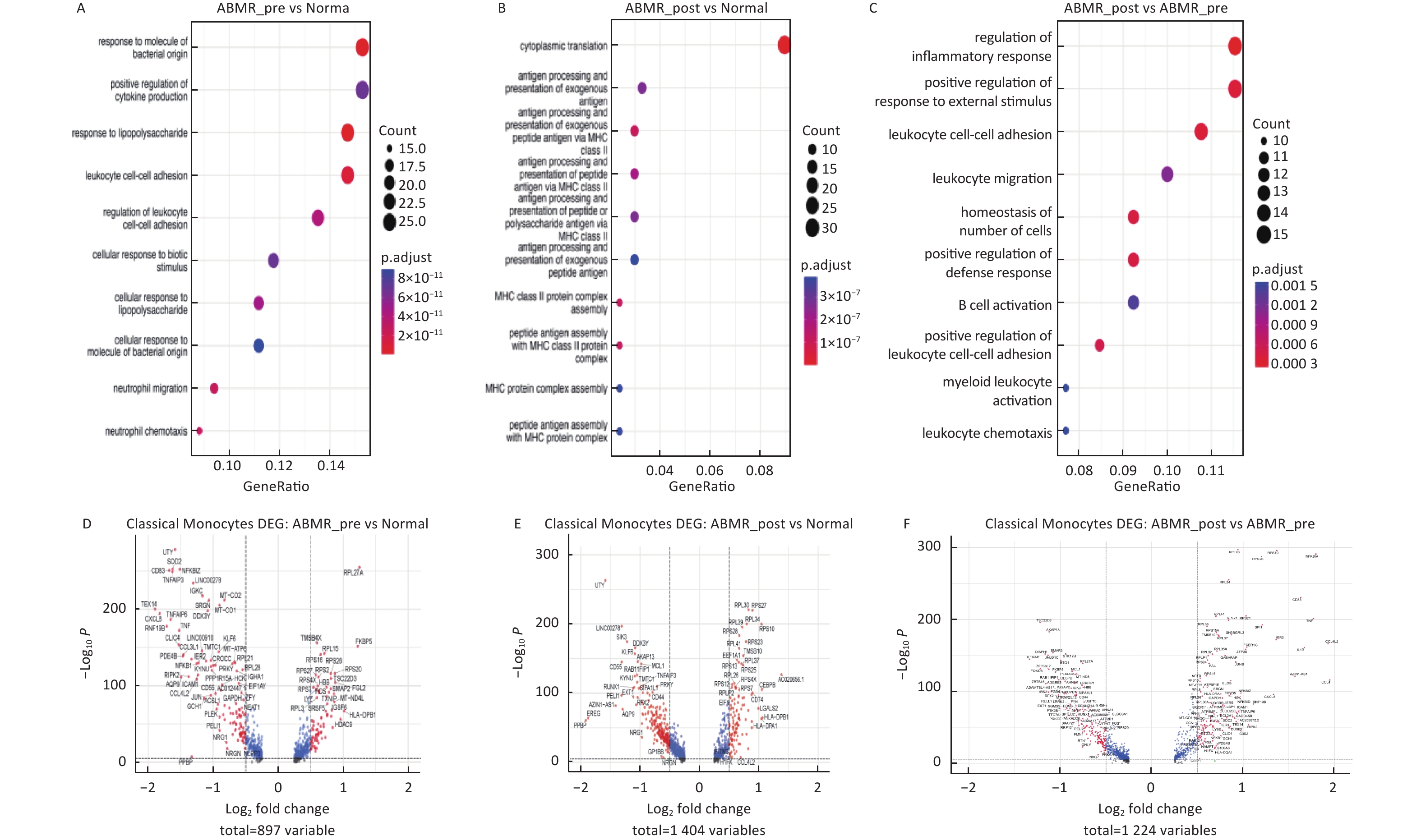
图 2
Fig.2 Differential gene and functional enrichment analysis in classical monocyte subsets A-C: signaling pathway enrichment analysis; D-F: differential gene analysis.
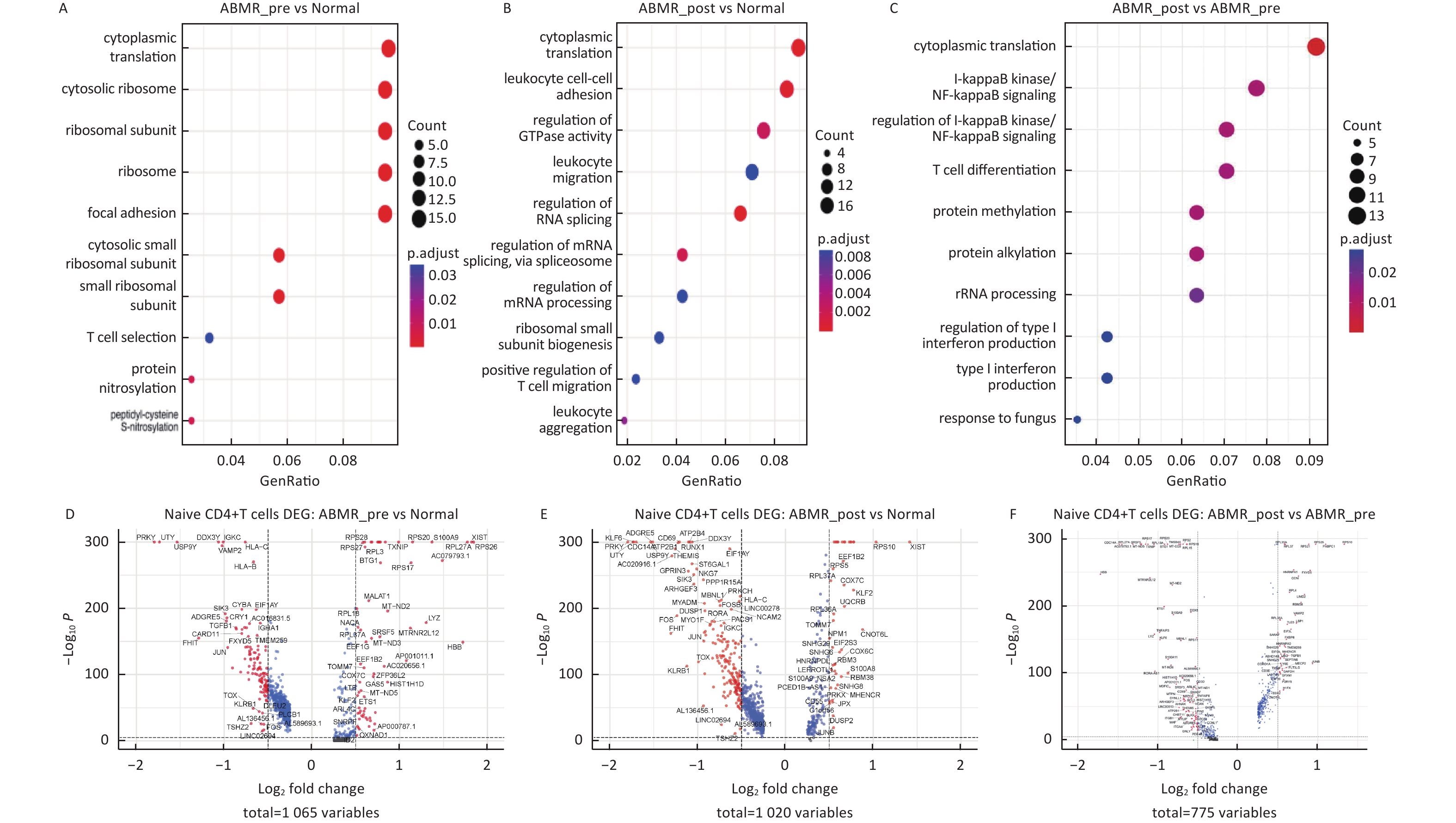
图 3
Fig.3 Differential gene expression, GO enrichment analysis in initial CD4+T cell subsets A-C: signaling pathway enrichment analysis; D-F: differential gene analysis.
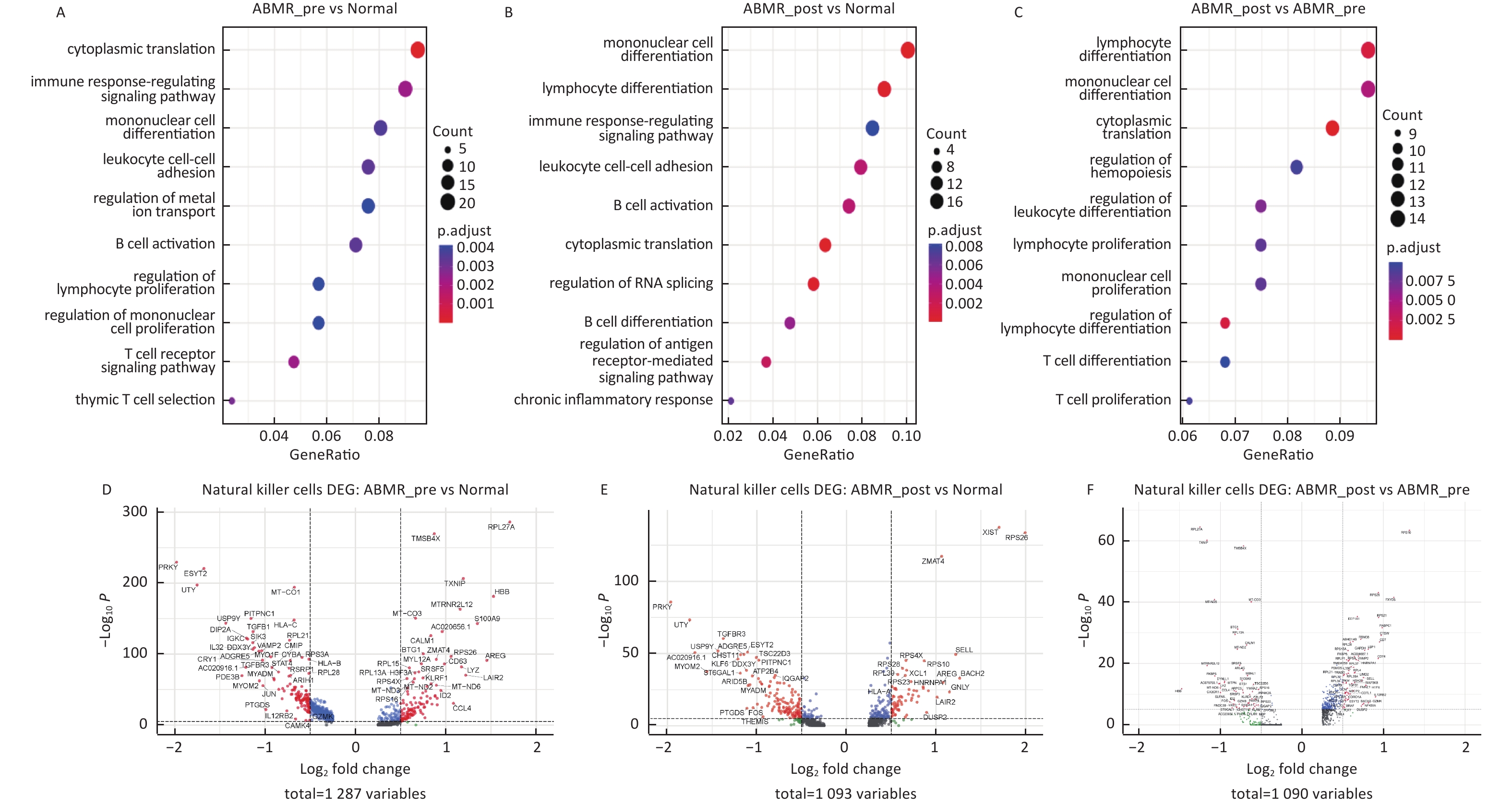
图 4
Fig.4 Differential gene expression, GO enrichment analysis in NK cell subsets A-C: signaling pathway enrichment analysis; D-F: differential gene analysis.
| Renal transplant recipient samples | CD83 (PC7-A) | CD52 (APC) |
| ABMR_pre (03-L1-F1-1) | ||
| ABMR_post (06-L H 17-C1) | ||
| Normal (04-W1-F1) |
表 3
Table 3 Geometric mean value of CD83 (PC7-A) and CD52 (APC) genes of for monocytes (n=3)
| Renal transplant recipient samples | CD83 (PC7-A) | CD52 (APC) |
| ABMR_pre (03-L1-F1-1) | ||
| ABMR_post (06-L H 17-C1) | ||
| Normal (04-W1-F1) |
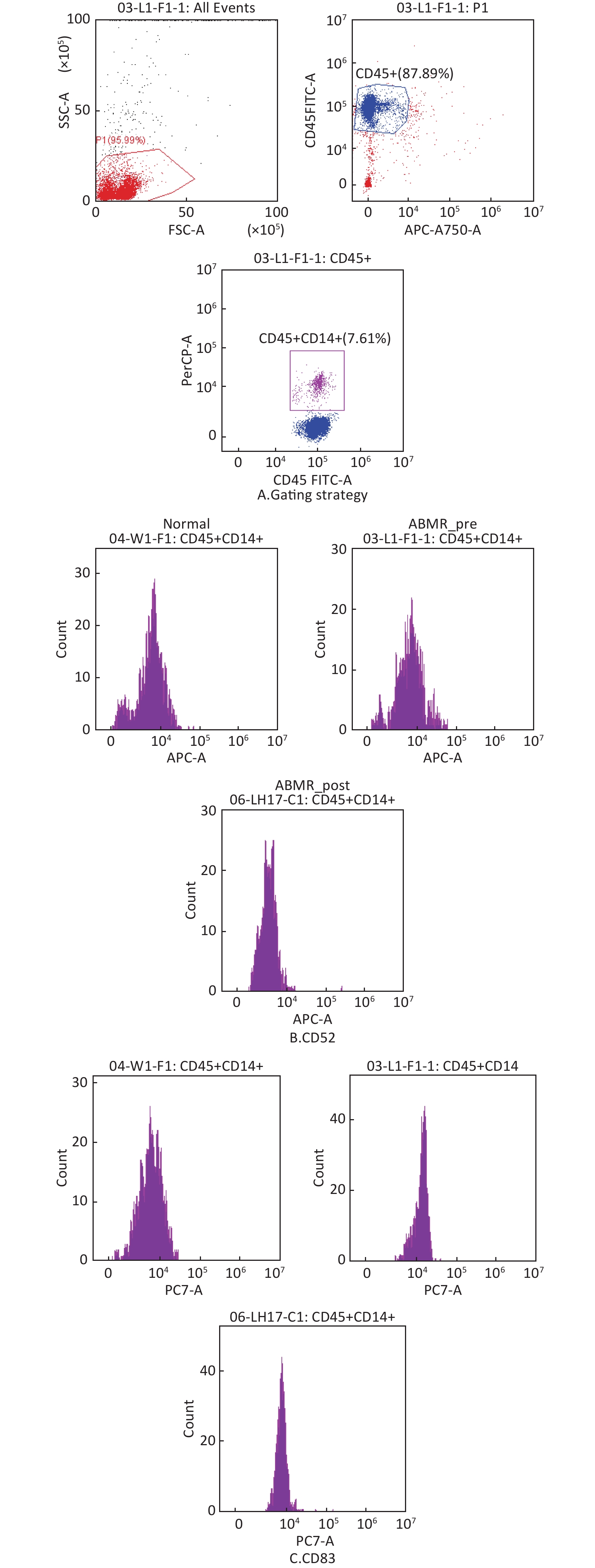
图 5
Fig.5 Flow cytometry to of changes in differentially expressed genes in classical monocytes of ABMR patients A: circle gate strategy for classical monocytes; B: histogram of CD52+; C: histogram of CD83+.
| Renal transplant recipient samples | CD69 (APC) |
| ABMR_pre (03-L1-F2-1) | |
| ABMR_post (06-L H 18-C2) | |
| Normal (04-W1-F2) |
表 4
Table 4 Geometric mean value of CD69 (APC) genes for naive CD4+T cells (n=3)
| Renal transplant recipient samples | CD69 (APC) |
| ABMR_pre (03-L1-F2-1) | |
| ABMR_post (06-L H 18-C2) | |
| Normal (04-W1-F2) |
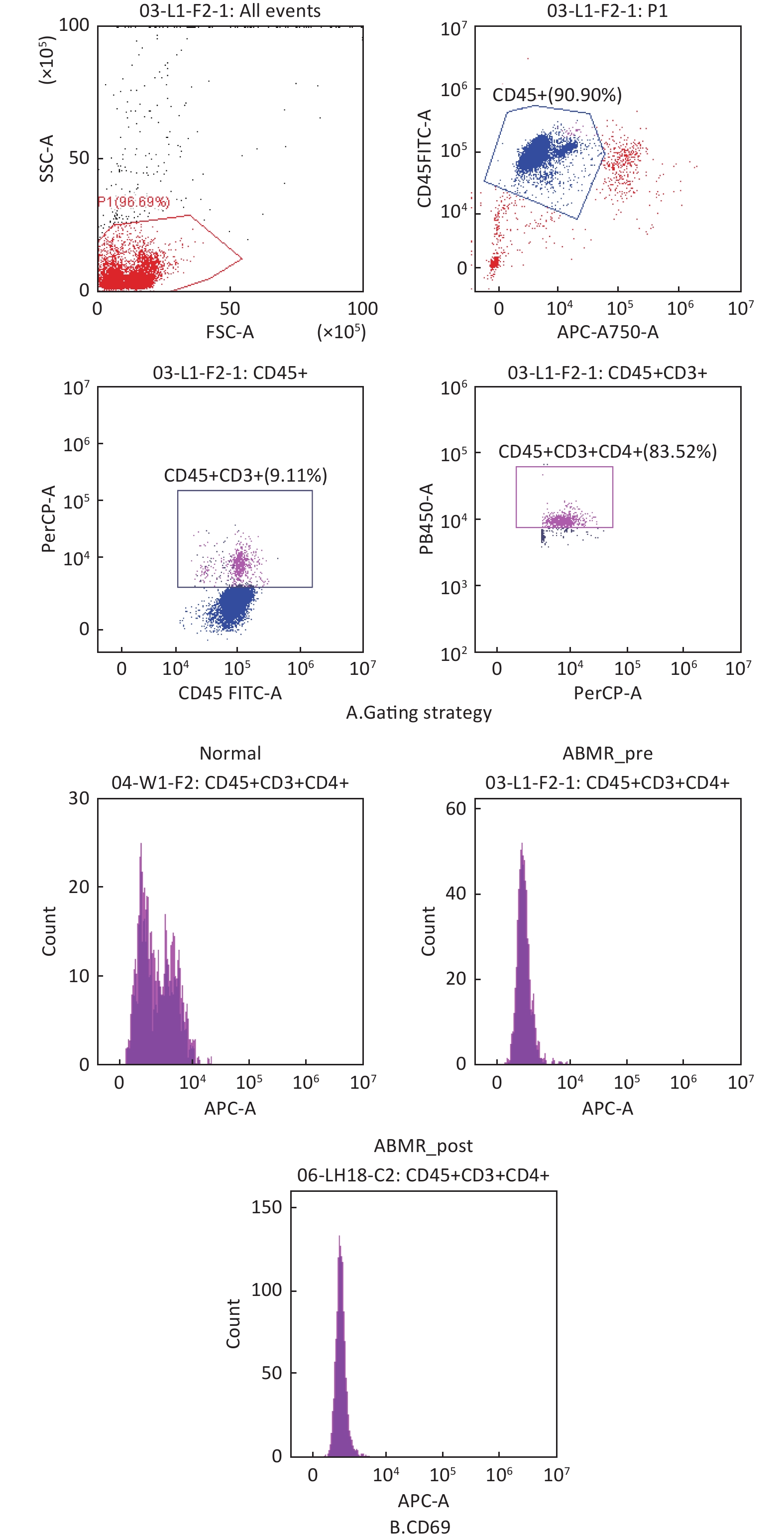
图 6 A: loop gate strategy for CD4+T cells; B: histogram of CD69+.
Fig.6 Flow cytometry validation of changes in differentially expressed genes in initial CD4+T cells from ABMR patients A: loop gate strategy for CD4+T cells; B: histogram of CD69+.
| Renal transplant recipient samples | CX3CR1 (PB450) | CD74 (APC) |
| ABMR_pre (03-L1-F3-1) | 472.5 | |
| ABMR_post (06-L H 19-C3) | 936.1 | 513.9 |
| Normal (07-GLB 19-C3) | 660.6 |
表 5
Table 5 Geometric mean value of CX3CR1 (PB450) and CD74 (APC) genes for NK cells
| Renal transplant recipient samples | CX3CR1 (PB450) | CD74 (APC) |
| ABMR_pre (03-L1-F3-1) | 472.5 | |
| ABMR_post (06-L H 19-C3) | 936.1 | 513.9 |
| Normal (07-GLB 19-C3) | 660.6 |
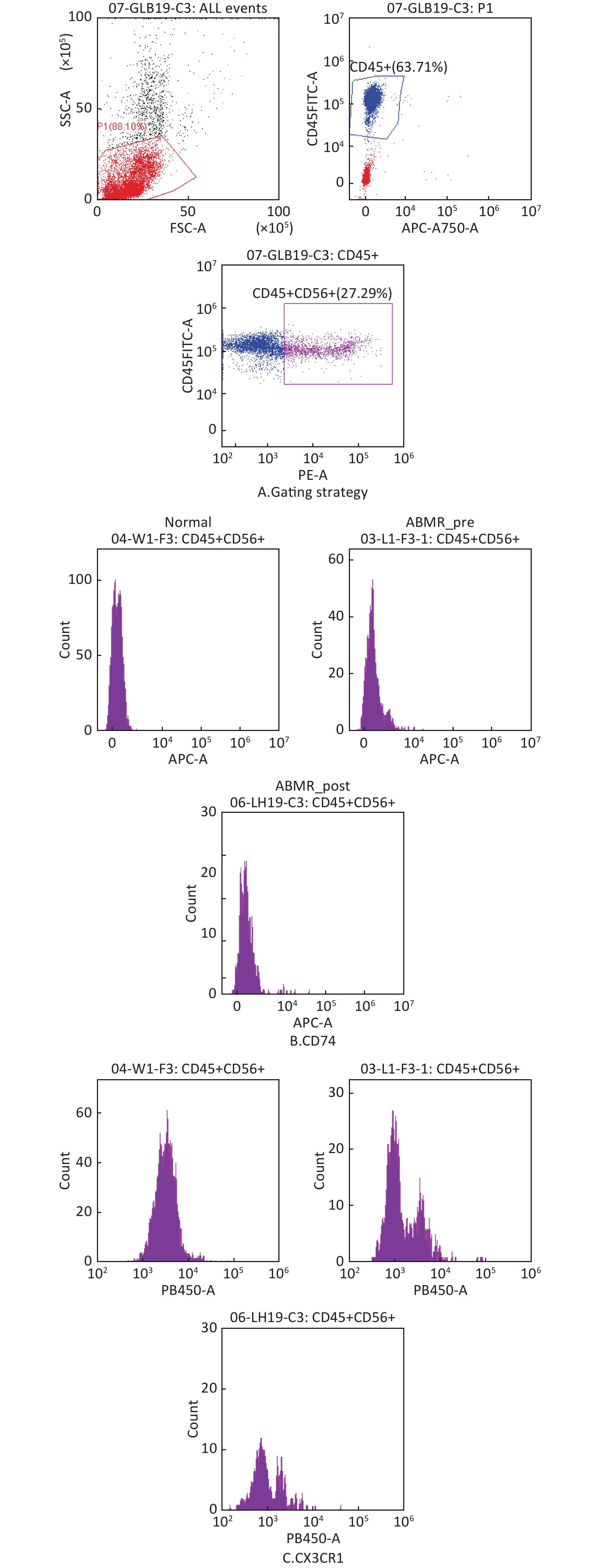
图 7
Fig.7 Flow cytometry validation of changes in differentially expressed genes in NK cells from ABMR patients (n=3) A: circle gate strategy of NK cells; B: histogram of CD74+; C: histogram of CX3CR1+.
| 1 |
Chen S, Zhou Y, Chen Y, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor[J]. Bioinformatics, 2018, 34 (17): i884- i890.
doi: 10.1093/bioinformatics/bty560 |
| 2 |
Leckie-Harre A, Silverman I, Wu H, et al. Sequencing of physically interacting cells in human kidney allograft rejection to infer contact-dependent immune cell transcription[J]. Transplantation, 2024, 108 (2): 421- 429.
doi: 10.1097/TP.0000000000004762 |
| 3 |
魏江浩, 窦古枫, 王振, 等. 肾移植术后慢性活动性抗体介导排斥反应的激素冲击治疗效果分析[J]. 山东医药, 2024, 64 (32): 16- 21.
doi: 10.3969/j.issn.1002-266X.2024.32.004 |
| 4 | 车福恒, 刘斌. 移植肾慢性活动性抗体介导排斥反应的研究进展[J]. 实用器官移植电子杂志, 2023, 11 (5): 489- 494. |
| 5 | Chen Y, Chen Y, Shi C, et al. Single-cell RNA sequencing reveals the immune landscape of human kidney allograft rejection[J]. J Am Soc Nephrol, 2020, 31 (11): 2745- 2760. |
| 6 |
Ni B, Yang C, Zhang J, et al. Rictor Ameliorates acute antibody-mediated rejection following kidney transplantation by suppressing macrophage M1 polarization through p65-NLRP3 axis[J]. Adv Sci (Weinh), 2025, 12 (34): e17119.
doi: 10.1002/advs.202417119 |
| 7 |
Haas M, Mirocha J, Huang E, et al. A banff-based histologic chronicity index is associated with graft loss in patients with a kidney transplant and antibody-mediated rejection[J]. Kidney Int, 2023, 103 (1): 187- 195.
doi: 10.1016/j.kint.2022.09.030 |
| 8 |
Shi W, Wan TT, Li HH, et al. Blockage of S100A8/A9 ameliorates septic nephropathy in mice[J]. Front Pharmacol, 2023, 14, 1172356.
doi: 10.3389/fphar.2023.1172356 |
| 9 |
Du L, Chen Y, Shi J, et al. Inhibition of S100A8/A9 ameliorates renal interstitial fibrosis in diabetic nephropathy[J]. Metabolism, 2023, 144, 155376.
doi: 10.1016/j.metabol.2022.155376 |
| 10 |
Pepper RJ, Wang HH, Rajakaruna GK, et al. S100A8/A9 (calprotectin) is critical for development of glomerulonephritis and promotes inflammatory leukocyte-renal cell interactions[J]. Am J Pathol, 2015, 185 (5): 1264- 1274.
doi: 10.1016/j.ajpath.2015.01.015 |
| 11 |
Li Y, Chen B, Yang X, et al. S100a8/a9 signaling causes mitochondrial dysfunction and cardiomyocyte death in response to ischemic/reperfusion injury[J]. Circulation, 2019, 140 (9): 751- 764.
doi: 10.1161/CIRCULATIONAHA.118.039262 |
| 12 |
Doberer K, Duerr M, Halloran PF, et al. A randomized clinical trial of anti-IL-6 antibody clazakizumab in late antibody-mediated kidney transplant rejection[J]. J Am Soc Nephrol, 2021, 32 (3): 708- 722.
doi: 10.1681/ASN.2020071106 |
| 13 |
Weerakoon H, Straube J, Lineburg K, et al. Expression of CD49f defines subsets of human regulatory T cells with divergent transcriptional landscape and function that correlate with ulcerative colitis disease activity[J]. Clin Transl Immunology, 2021, 10 (9): e1334.
doi: 10.1002/cti2.1334 |
| 14 |
Lu J, Wang W, Li P, et al. miR-146a regulates regulatory T cells to suppress heart transplant rejection in mice[J]. Cell Death Discov, 2021, 7 (1): 165.
doi: 10.1038/s41420-021-00534-9 |
| 15 | Schürch CM, Bhattacharya N, Barbash O, et al. Single-cell RNA sequencing reveals the cellular architecture of chronic kidney allograft rejection[J]. Nat Med, 2020, 26 (2): 211- 221. |
| 16 | Li H, Kaminski MS, Li N, et al. Single-cell profiling of human kidney allografts reveals an immunoregulatory role for NK cells in acute rejection[J]. J Clin Invest, 2019, 129 (10): 4174- 4189. |
| 17 |
Zhao Y, Su H, Shen X, et al. The immunological function of CD52 and its targeting in organ transplantation[J]. Inflamm Res, 2017, 66 (7): 571- 578.
doi: 10.1007/s00011-017-1032-8 |
| 18 |
Borghese F, Clanchy FI. CD74: an emerging opportunity as a therapeutic target in cancer and autoimmune disease[J]. Expert Opin Ther Targets, 2011, 15 (3): 237- 251.
doi: 10.1517/14728222.2011.550879 |
| 19 |
Cormican S, Griffin MD. Fractalkine (CX3CL1) and its receptor CX3CR1: A promising therapeutic target in chronic kidney disease ?[J]. Front Immunol, 2021, 12, 664202.
doi: 10.3389/fimmu.2021.664202 |
| 20 |
Li X, Li S, Wu B, et al. Landscape of immune cells heterogeneity in liver transplantation by single-cell RNA sequencing analysis[J]. Front Immunol, 2022, 13, 890019.
doi: 10.3389/fimmu.2022.890019 |
| 21 |
Xia P, Ji X, Yan L, et al. Roles of S100A8, S100A9 and S100A12 in infection, inflammation and immunity[J]. Immunology, 2024, 171 (3): 365- 376.
doi: 10.1111/imm.13722 |
| 22 |
Bui TM, Wiesolek HL, Sumagin R. ICAM-1: a master regulator of cellular responses in inflammation, injury resolution, and tumorigenesis[J]. J Leukoc Biol, 2020, 108 (3): 787- 799.
doi: 10.1002/JLB.2MR0220-549R |
| 23 |
Blanco-Domínguez R, de la Fuente H, Rodríguez C, et al. CD69 expression on regulatory T cells protects from immune damage after myocardial infarction[J]. J Clin Invest, 2022, 132 (21): e152418.
doi: 10.1172/JCI152418 |
| 24 |
Liu J, Yue WL, Fan HZ, et al. Correlation of cTfh cells and memory B cells with AMR after renal transplantation[J]. Transpl Immunol, 2024, 86, 102095.
doi: 10.1016/j.trim.2024.102095 |
| 25 |
Li Z, Ju X, Silveira PA, et al. CD83: activation marker for antigen presenting cells and its therapeutic potential[J]. Front Immunol, 2019, 10, 1312.
doi: 10.3389/fimmu.2019.01312 |
| 26 |
Chukwu CA, Spiers HVM, Middleton R, et al. Alemtuzumab in renal transplantation. Reviews of literature and usage in the United Kingdom[J]. Transplant Rev (Orlando), 2022, 36 (2): 100686.
doi: 10.1016/j.trre.2022.100686 |
| 27 | Kuppe C, Ibrahim MM, Kranz J, et al. Spatial single-cell analysis identifies an interstitial immune network in human kidney allografts[J]. Nature, 2021, 593 (7858): 253- 258. |
| 28 |
Serrano OK, Friedmann P, Ahsanuddin S, et al. Outcomes associated with steroid avoidance and alemtuzumab among kidney transplant recipients[J]. Clin J Am Soc Nephrol, 2015, 10 (11): 2030- 2038.
doi: 10.2215/CJN.12161214 |
| 29 |
van der Zwan M, Baan CC, van Gelder T, et al. Review of the clinical pharmacokinetics and pharmacodynamics of alemtuzumab and its use in kidney transplantation[J]. Clin Pharmacokinet, 2018, 57 (2): 191- 207.
doi: 10.1007/s40262-017-0573-x |
| 30 |
Yagisawa T, Tanaka T, Miyairi S, et al. In the absence of natural killer cell activation donor-specific antibody mediates chronic, but not acute, kidney allograft rejection[J]. Kidney Int, 2019, 95 (2): 350- 336.
doi: 10.1016/j.kint.2018.08.041 |
| [1] | 陈冰, 左笑丛, 李新刚, 尚德为, 周佩军, 丁俊杰, 相小强, 邱晓燕, 王卓, 李晓宇, 张弋, 赵维, 王玉珠, 高建军, 焦正. 模型引导的抗排异治疗患者他克莫司精准用药专家共识[J]. 中国临床药理学与治疗学, 2025, 30(4): 433-445. |
| [2] | 常钊, 周宇雪, 张胜男, 吕萌. 儿童肾病综合征患者他克莫司血药浓度变异性与临床疗效关系评价及影响因素分析[J]. 中国临床药理学与治疗学, 2025, 30(11): 1524-1529. |
| [3] | 王学彬, 高丽红, 张凌鹏, 杨云云, 刘红霞, 王卓, 孙华君. 他克莫司治疗药物监测异常结果干预及评价研究[J]. 中国临床药理学与治疗学, 2024, 29(12): 1322-1328. |
| [4] | 柴玉慧, 杨云云, 权丹妮, 张凌鹏, 高丽红, 王学彬, 王卓. miR-144-5p调控的FoxO1作为肾移植抗体介导排斥反应潜在治疗靶点的生物信息学分析[J]. 中国临床药理学与治疗学, 2024, 29(12): 1359-1366. |
| [5] | 李 坤, 李璐璐, 李楠楠, 胡卫红, 周建超. 成人肾移植术后血糖控制和CYP3A5基因多态性对他克莫司谷浓度的影响[J]. 中国临床药理学与治疗学, 2023, 28(7): 767-774. |
| [6] | 陈以宁, 肖 云, 韩晓雨, 黄璐璐, 马景胜, 熊寒斌, 傅 毓, 汪保林, 郭晓晖, 钟 林, 敖检根, 何佳珂. 肝移植术后排斥合并高血糖的诊治和用药的实践与思考[J]. 中国临床药理学与治疗学, 2023, 28(5): 550-555. |
| [7] | 王爽, 刘嘉, 张月丽. POR*28基因多态性与中国肾移植患者他克莫司稳定剂量的相关性研究[J]. 中国临床药理学与治疗学, 2022, 27(9): 991-997. |
| [8] | 杜文鹏, 敖检根, 陶 祎, 吴观胜, 何佳珂. 自身免疫疾病患者他克莫司稳态血药浓度影响因素的研究[J]. 中国临床药理学与治疗学, 2022, 27(6): 645-651. |
| [9] | 吴懿, 方芳, 陈瑛, 樊军卫. 供受体CYP3A5基因型对肝移植术后早期他克莫司谷浓度的影响作用及其临床意义[J]. 中国临床药理学与治疗学, 2021, 26(6): 631-639. |
| [10] | 刘建辉, 周根鸿, 李春林, 鲍美华, 刘方怡. 生物信息学筛选骨髓增生异常综合征相关长链非编码RNA[J]. 中国临床药理学与治疗学, 2020, 25(12): 1351-1356. |
| [11] | 吴懿, 王让让, 王兆文, 樊军卫. 受体SLCO1B1 rs2291075位点多态性对肝移植术后早期他克莫司剂量校正谷浓度的影响[J]. 中国临床药理学与治疗学, 2020, 25(11): 1268-1275. |
| [12] | 闫文龙, 疏树华, 李娟. 5%碳酸氢钠严格控制代谢性酸中毒对活体肾移植患者术后早期肾功能的影响[J]. 中国临床药理学与治疗学, 2020, 25(10): 1139-1144. |
| [13] | 刘宇佳,李 莉,胡晓平,钟里科,李晶晶,黄 萍,张轶雯. 基于生物信息学的胃癌特征基因网络关键节点及预后关联分析[J]. 中国临床药理学与治疗学, 2019, 24(8): 852-859. |
| [14] | 方 茜,屈 强,肖湘成. 他克莫司治疗特发性膜性肾病研究进展[J]. 中国临床药理学与治疗学, 2019, 24(2): 228-234. |
| [15] | 张轶雯,楼倩雯,钟里科,潘宗富,方 罗,黄 萍. 基于GEO数据库的近端与远端结肠癌特征基因生物信息学研究[J]. 中国临床药理学与治疗学, 2018, 23(4): 422-427. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||