| 1 |
Chen S, Zhou Y, Chen Y, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor[J]. Bioinformatics, 2018, 34 (17): i884- i890.
doi: 10.1093/bioinformatics/bty560
|
| 2 |
Leckie-Harre A, Silverman I, Wu H, et al. Sequencing of physically interacting cells in human kidney allograft rejection to infer contact-dependent immune cell transcription[J]. Transplantation, 2024, 108 (2): 421- 429.
doi: 10.1097/TP.0000000000004762
|
| 3 |
魏江浩, 窦古枫, 王振, 等. 肾移植术后慢性活动性抗体介导排斥反应的激素冲击治疗效果分析[J]. 山东医药, 2024, 64 (32): 16- 21.
doi: 10.3969/j.issn.1002-266X.2024.32.004
|
| 4 |
车福恒, 刘斌. 移植肾慢性活动性抗体介导排斥反应的研究进展[J]. 实用器官移植电子杂志, 2023, 11 (5): 489- 494.
|
| 5 |
Chen Y, Chen Y, Shi C, et al. Single-cell RNA sequencing reveals the immune landscape of human kidney allograft rejection[J]. J Am Soc Nephrol, 2020, 31 (11): 2745- 2760.
|
| 6 |
Ni B, Yang C, Zhang J, et al. Rictor Ameliorates acute antibody-mediated rejection following kidney transplantation by suppressing macrophage M1 polarization through p65-NLRP3 axis[J]. Adv Sci (Weinh), 2025, 12 (34): e17119.
doi: 10.1002/advs.202417119
|
| 7 |
Haas M, Mirocha J, Huang E, et al. A banff-based histologic chronicity index is associated with graft loss in patients with a kidney transplant and antibody-mediated rejection[J]. Kidney Int, 2023, 103 (1): 187- 195.
doi: 10.1016/j.kint.2022.09.030
|
| 8 |
Shi W, Wan TT, Li HH, et al. Blockage of S100A8/A9 ameliorates septic nephropathy in mice[J]. Front Pharmacol, 2023, 14, 1172356.
doi: 10.3389/fphar.2023.1172356
|
| 9 |
Du L, Chen Y, Shi J, et al. Inhibition of S100A8/A9 ameliorates renal interstitial fibrosis in diabetic nephropathy[J]. Metabolism, 2023, 144, 155376.
doi: 10.1016/j.metabol.2022.155376
|
| 10 |
Pepper RJ, Wang HH, Rajakaruna GK, et al. S100A8/A9 (calprotectin) is critical for development of glomerulonephritis and promotes inflammatory leukocyte-renal cell interactions[J]. Am J Pathol, 2015, 185 (5): 1264- 1274.
doi: 10.1016/j.ajpath.2015.01.015
|
| 11 |
Li Y, Chen B, Yang X, et al. S100a8/a9 signaling causes mitochondrial dysfunction and cardiomyocyte death in response to ischemic/reperfusion injury[J]. Circulation, 2019, 140 (9): 751- 764.
doi: 10.1161/CIRCULATIONAHA.118.039262
|
| 12 |
Doberer K, Duerr M, Halloran PF, et al. A randomized clinical trial of anti-IL-6 antibody clazakizumab in late antibody-mediated kidney transplant rejection[J]. J Am Soc Nephrol, 2021, 32 (3): 708- 722.
doi: 10.1681/ASN.2020071106
|
| 13 |
Weerakoon H, Straube J, Lineburg K, et al. Expression of CD49f defines subsets of human regulatory T cells with divergent transcriptional landscape and function that correlate with ulcerative colitis disease activity[J]. Clin Transl Immunology, 2021, 10 (9): e1334.
doi: 10.1002/cti2.1334
|
| 14 |
Lu J, Wang W, Li P, et al. miR-146a regulates regulatory T cells to suppress heart transplant rejection in mice[J]. Cell Death Discov, 2021, 7 (1): 165.
doi: 10.1038/s41420-021-00534-9
|
| 15 |
Schürch CM, Bhattacharya N, Barbash O, et al. Single-cell RNA sequencing reveals the cellular architecture of chronic kidney allograft rejection[J]. Nat Med, 2020, 26 (2): 211- 221.
|
| 16 |
Li H, Kaminski MS, Li N, et al. Single-cell profiling of human kidney allografts reveals an immunoregulatory role for NK cells in acute rejection[J]. J Clin Invest, 2019, 129 (10): 4174- 4189.
|
| 17 |
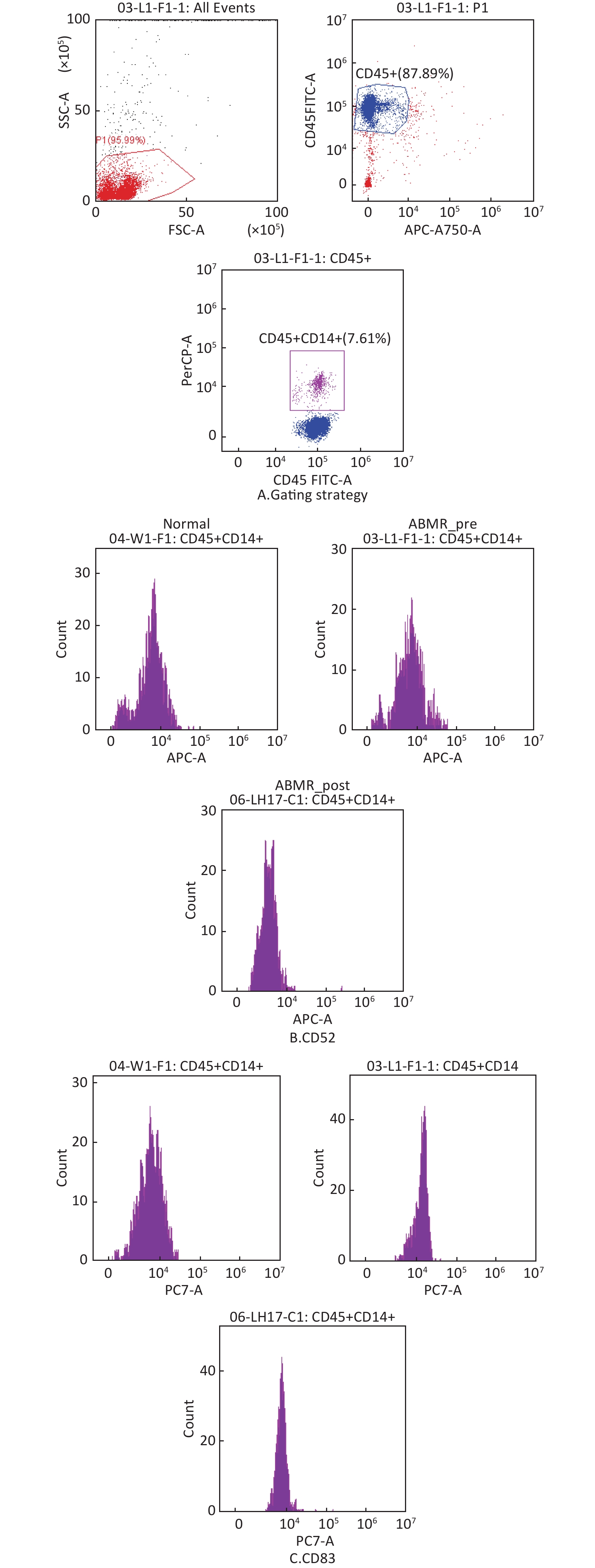
Zhao Y, Su H, Shen X, et al. The immunological function of CD52 and its targeting in organ transplantation[J]. Inflamm Res, 2017, 66 (7): 571- 578.
doi: 10.1007/s00011-017-1032-8
|
| 18 |
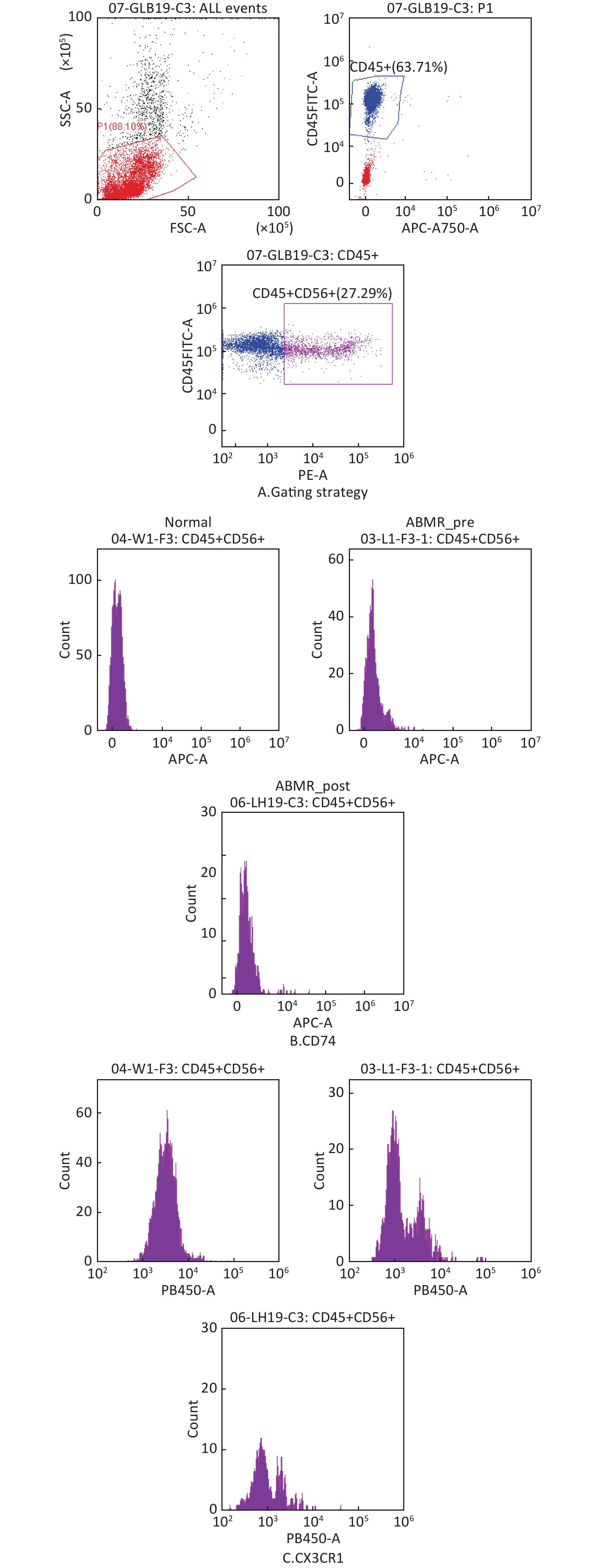
Borghese F, Clanchy FI. CD74: an emerging opportunity as a therapeutic target in cancer and autoimmune disease[J]. Expert Opin Ther Targets, 2011, 15 (3): 237- 251.
doi: 10.1517/14728222.2011.550879
|
| 19 |
Cormican S, Griffin MD. Fractalkine (CX3CL1) and its receptor CX3CR1: A promising therapeutic target in chronic kidney disease ?[J]. Front Immunol, 2021, 12, 664202.
doi: 10.3389/fimmu.2021.664202
|
| 20 |
Li X, Li S, Wu B, et al. Landscape of immune cells heterogeneity in liver transplantation by single-cell RNA sequencing analysis[J]. Front Immunol, 2022, 13, 890019.
doi: 10.3389/fimmu.2022.890019
|
| 21 |
Xia P, Ji X, Yan L, et al. Roles of S100A8, S100A9 and S100A12 in infection, inflammation and immunity[J]. Immunology, 2024, 171 (3): 365- 376.
doi: 10.1111/imm.13722
|
| 22 |
Bui TM, Wiesolek HL, Sumagin R. ICAM-1: a master regulator of cellular responses in inflammation, injury resolution, and tumorigenesis[J]. J Leukoc Biol, 2020, 108 (3): 787- 799.
doi: 10.1002/JLB.2MR0220-549R
|
| 23 |
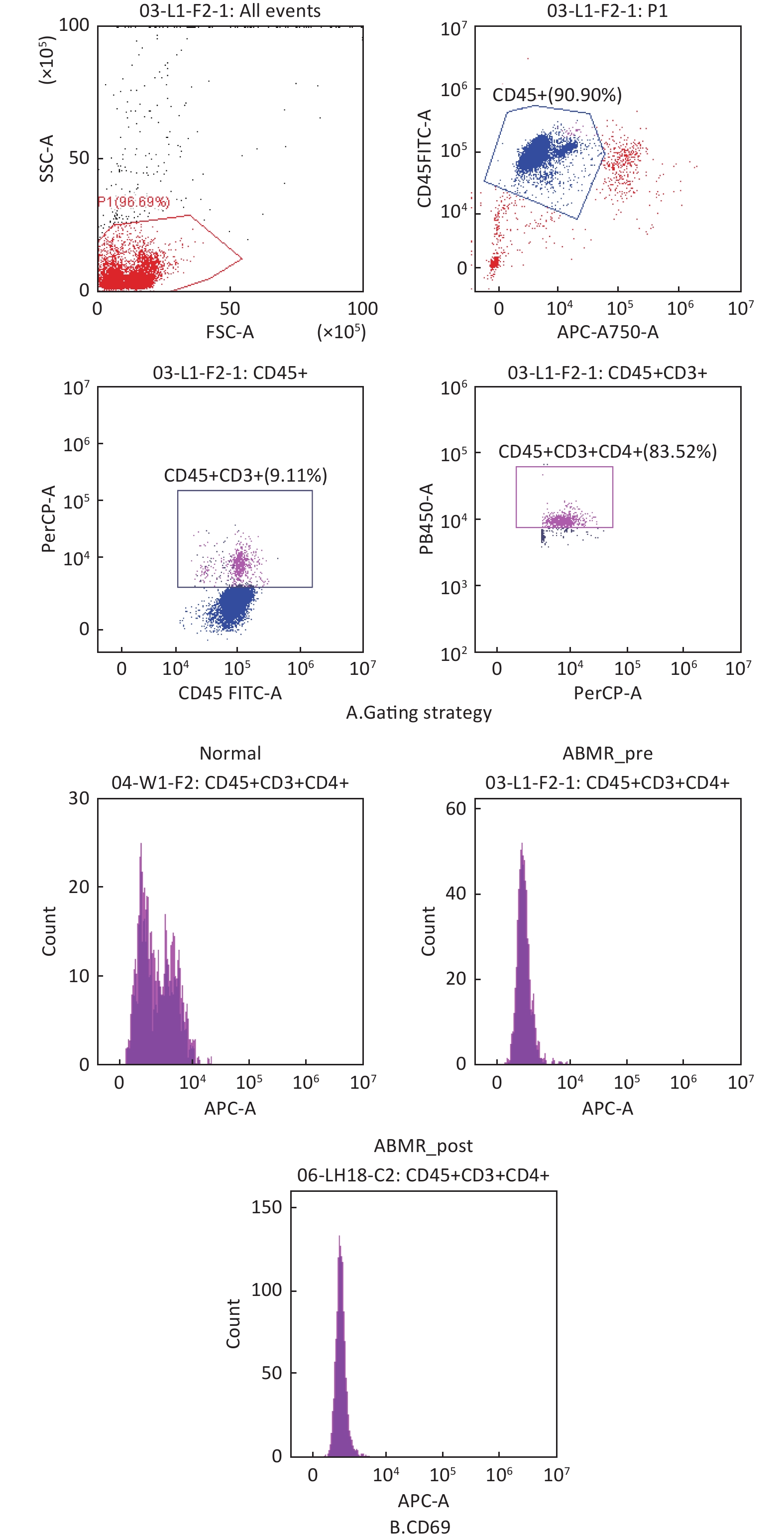
Blanco-Domínguez R, de la Fuente H, Rodríguez C, et al. CD69 expression on regulatory T cells protects from immune damage after myocardial infarction[J]. J Clin Invest, 2022, 132 (21): e152418.
doi: 10.1172/JCI152418
|
| 24 |
Liu J, Yue WL, Fan HZ, et al. Correlation of cTfh cells and memory B cells with AMR after renal transplantation[J]. Transpl Immunol, 2024, 86, 102095.
doi: 10.1016/j.trim.2024.102095
|
| 25 |
Li Z, Ju X, Silveira PA, et al. CD83: activation marker for antigen presenting cells and its therapeutic potential[J]. Front Immunol, 2019, 10, 1312.
doi: 10.3389/fimmu.2019.01312
|
| 26 |
Chukwu CA, Spiers HVM, Middleton R, et al. Alemtuzumab in renal transplantation. Reviews of literature and usage in the United Kingdom[J]. Transplant Rev (Orlando), 2022, 36 (2): 100686.
doi: 10.1016/j.trre.2022.100686
|
| 27 |
Kuppe C, Ibrahim MM, Kranz J, et al. Spatial single-cell analysis identifies an interstitial immune network in human kidney allografts[J]. Nature, 2021, 593 (7858): 253- 258.
|
| 28 |
Serrano OK, Friedmann P, Ahsanuddin S, et al. Outcomes associated with steroid avoidance and alemtuzumab among kidney transplant recipients[J]. Clin J Am Soc Nephrol, 2015, 10 (11): 2030- 2038.
doi: 10.2215/CJN.12161214
|
| 29 |
van der Zwan M, Baan CC, van Gelder T, et al. Review of the clinical pharmacokinetics and pharmacodynamics of alemtuzumab and its use in kidney transplantation[J]. Clin Pharmacokinet, 2018, 57 (2): 191- 207.
doi: 10.1007/s40262-017-0573-x
|
| 30 |
Yagisawa T, Tanaka T, Miyairi S, et al. In the absence of natural killer cell activation donor-specific antibody mediates chronic, but not acute, kidney allograft rejection[J]. Kidney Int, 2019, 95 (2): 350- 336.
doi: 10.1016/j.kint.2018.08.041
|